Finding, exploring & reusing published projects
Finding projects
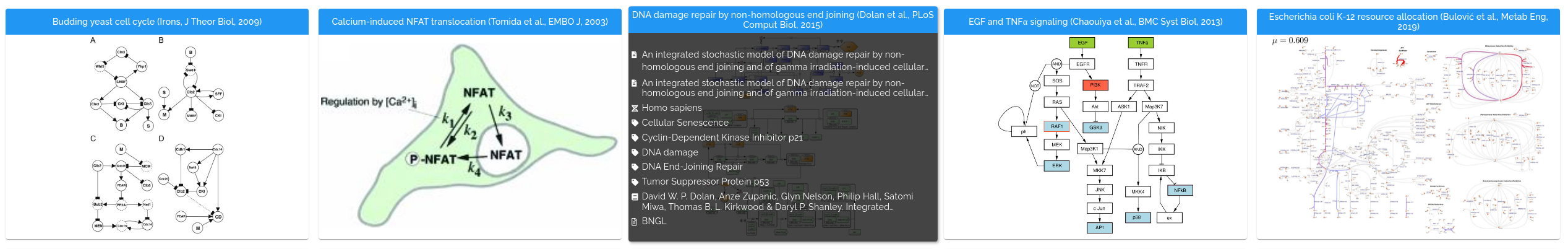
Published projects can be browsed at https://biosimulations.org/projects. Each card presents a project, with a thumbnail and title. Mousing over the thumbnail shows additional details about the project. You can customize the attributes that are displayed.

Selecting attributes
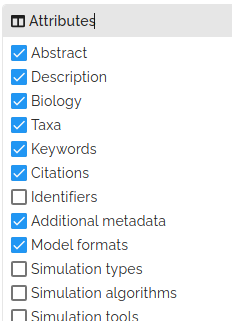 Clicking on the search icon in the top-right corner of the page opens a menu with an attributes sub-menu. From here, you can select the attributes that are displayed. Selecting a field will add the attributes to the details presented in the project card when you mouse over each thumbnail.
Clicking on the search icon in the top-right corner of the page opens a menu with an attributes sub-menu. From here, you can select the attributes that are displayed. Selecting a field will add the attributes to the details presented in the project card when you mouse over each thumbnail.
Searching for projects
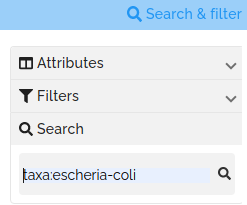 Clicking on the the search icon at the top right of the page opens a search box. A search term, such as 'metabolism' can be entered in the search box. By default, the search term is searched against each attribute of each project. Optionally, you can restrict the search to specific attributes. For example, if you want to search for projects that have the taxa 'Escherichia coli', you can enter 'taxa:Escherichia coli' in the search box. For attributes with spaces in the name, replace these spaces with "-". For example, the term "last-updated:2020" can be used to search for projects that contain the value "2020" in the ;last updated' attribute. A list of the available search fields is available in the FAQs.
Clicking on the the search icon at the top right of the page opens a search box. A search term, such as 'metabolism' can be entered in the search box. By default, the search term is searched against each attribute of each project. Optionally, you can restrict the search to specific attributes. For example, if you want to search for projects that have the taxa 'Escherichia coli', you can enter 'taxa:Escherichia coli' in the search box. For attributes with spaces in the name, replace these spaces with "-". For example, the term "last-updated:2020" can be used to search for projects that contain the value "2020" in the ;last updated' attribute. A list of the available search fields is available in the FAQs.
Filtering projects
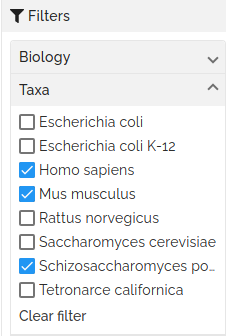 The list of displayed projects can be filtered by the values of their attributes. For each available attribute, a menu of values is presented. Selecting a value will filter the list of projects to include only those with the selected values.
The list of displayed projects can be filtered by the values of their attributes. For each available attribute, a menu of values is presented. Selecting a value will filter the list of projects to include only those with the selected values.
Exploring projects
Clicking on a project card opens a page with the project details. The "Overview" tab provides metadata about the selected project, as well as information about underlying model and simulation run. The "Select chart" tab allows you to configure visualizations of the simulation results that can then be viewed on the "View chart" tab. The "Files" tab provides downloads for the files of the project.
Metadata
BioSimulations collects metadata to enable searching, browsing and discovering projects. The metadata includes information about authorship, license, funding and other provenance information. It also includes information about the modelled system, such as the modelled organism, and tags that describe the project.

Visualizations
The results of simulations can be visualized using both predefined and custom visualizations. The "Select chart" tab allows you to select from pre-defined visualizations that the author included, including both basic charts described with SED-ML and more complex visualizations described with Vega. Additionally, you can create your own custom visualizations by selecting one of the "Design a chart" options including histograms, heatmaps and lineplots. Selecting a plot time will open an additional menu with configuration options to select the datasets to be plotted.
Once you have configured your visualization, you can view it by clicking on the "View chart" button. The "Export to Vega" button will export the visualization to a Vega specification, which enables greater user customization of the visual. More information for using Vega with BioSimulations in available here.
More information about creating SED-ML and Vega visualizations is available here and here.
Simulation runs
 More detailed information about the execution of the project and its results can be viewed by following the links to the runBioSimulations page for the project. The "Logs" tab provides detailed output of the simulation execution, including each individual simulation task and the outputs (reports and plots) produced by the simulation. Each task of the simulation is presented as a collapsible section that can be expanded to show the outputs. Both structured log files and raw output files can be downloaded from the links.
More detailed information about the execution of the project and its results can be viewed by following the links to the runBioSimulations page for the project. The "Logs" tab provides detailed output of the simulation execution, including each individual simulation task and the outputs (reports and plots) produced by the simulation. Each task of the simulation is presented as a collapsible section that can be expanded to show the outputs. Both structured log files and raw output files can be downloaded from the links.
Reusing projects
Creating and executing variants of simulations with runBioSimulations
In addition to this full-featured web application, runBioSimulations provides a simpler web application and REST API for executing simulations. runBioSimulations simply enables users to execute COMBINE/OMEX archives using a variety of simulation tools and generate time series plots of their results. runBioSimulations does not require an account.
Downloading projects and executing them with your own computers
Downloading projects
The models, simulations, and visualizations in BioSimulations can be programmatically obtained using our REST API. Documentation for the API is available at the same URL.
Recommended tools for further exploring simulation projects
BioSimulations provides basic capabilities for reproducing and reusing a wide range of biomodeling projects. For further work, we encourage users to use the domain-specific online platforms, desktop programs, and libraries outlined below. Consistent interfaces to the desktop and library tools below are available from BioSimulators, including Docker images, command-line interfaces and Python APIs. More information about obtaining and using these tools is available from BioSimulators.
Warning
While the BioSimulators interfaces to these tools support SED-ML and the COMBINE/OMEX archive format, the primary versions of most of the tools below do not support these formats or do not support them consistently with the specifications of the SED-ML format.
| Framework | Language | Online programs | Desktop programs | Libraries |
|---|---|---|---|---|
| Continuous kinetic | BNGL | BioNetGen | pyBioNetGen | |
| Continuous kinetic | CellML | OpenCOR | OpenCOR | |
| Continuous kinetic | NeuroML | NetPyNe, NEURON, pyNeuroML | NetPyNe, NEURON, pyNeuroML | |
| Continuous kinetic | SBML | JWS Online | BioNetGen, COPASI, tellurium, VCell | AMICI, GillesPy2, libRoadRunner, LibSBMLSim, pyBioNetGen, PySCeS |
| Continuous kinetic | XPP ODE | XPP | ||
| Discrete kinetic | BNGL | BioNetGen | pyBioNetGen | |
| Discrete kinetic | SBML | StochSS | BioNetGen, COPASI, tellurium, VCell | GillesPy2, libRoadRunner, pyBioNetGen |
| Flux balance | SBML | Fluxer | CBMPy | CBMPy, COBRApy |
| Logical | GINsim | GINsim | ||
| Logical | SBML | Cell Collective | GINsim | BoolNet |
| MASS | SBML | MASSpy | ||
| Resource balance | RBA XML | RBApy | ||
| Spatial discrete | Smoldyn | Smoldyn | Smoldyn |